But last year I did a stint in a computational lab, so somewhat reluctantly I learned Shell scripting. (Shell scripting is like Sesame-Street-level computer programming.) The other day I found myself needing to search a large mass of protein sequence for a motif. How to do it? The Shell command grep.
Here’s the thing. If you have a Macintosh, then grep is a super-pimped out search feature that is inside your computer right now. For searching inside large text files, it’s way more powerful than Spotlight or whatever they’re calling that magnifying glass in the corner these days. You’ll need to know some kindergarten-level computer programming to be able to use it, but it’s totally worthwhile.
Start by taking the stuff you want to search and pasting it into a text file. It’s important that it’s plain text and that there aren’t any spaces in the file name. Save and quit.
Open the computer program Terminal. It’s in Applications > Utilities.
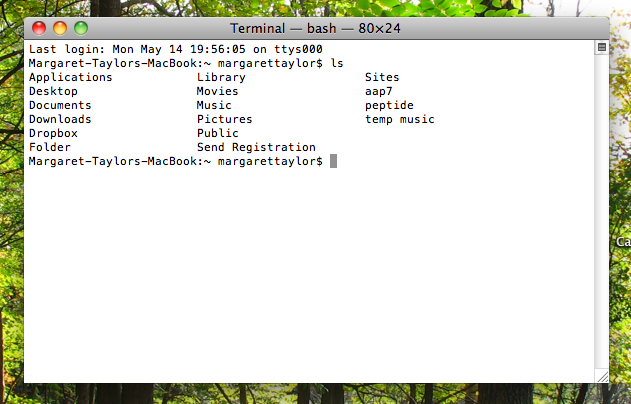
Type ls and hit return. That gives you a top-level list of all the folders on your computer. Type cd and the name of a folder to open it. Keep going until you’ve opened the folder that has your file in it. (Your folders had better not have spaces in their names!)
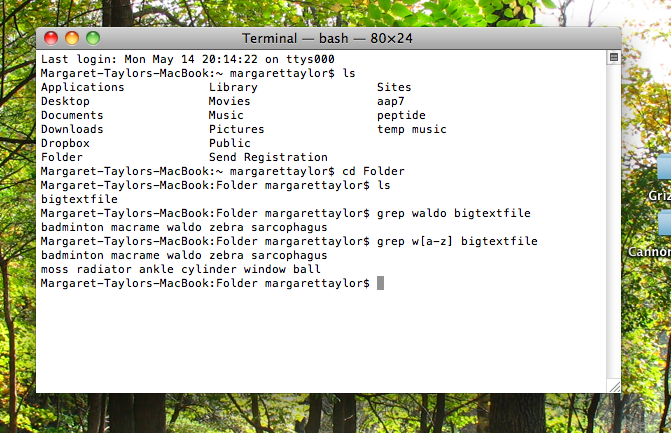
Type grep whatyou'relookingfor nameoftextfile.
But can’t ⌘F do the same thing just as well? Ah, here is where grep is a pimped-out search feature. It can use regular expressions. Here’s a longer explanation of regular expressions, but in short, they let you specify any sort of search criteria you could possibly imagine. Here’s a real simple example where I remember that the word I’m searching for starts with w, but I don’t know what comes next:
And that’s the magic of grep!
* “Principal component analysis (PCA) is a mathematical procedure that uses an orthogonal transformation to convert a set of observations of possibly correlated variables into a set of values of linearly uncorrelated variables called principal components,” says Wikipedia.